Research: Comparing Cellular Cousins
motivation
Basically I wanted to compare naked mole rat fibroblasts and human induced pluripotent stem cells against human fibroblasts to try to find genes that prevent aging, since naked mole rats and IPSCs don’t age while human cells do. The general metric would be: abs[(NMRCell - humanCell) + (humanIPSC - humanCell)]/abs(NMRCell - humanIPSC), so that hits are rewarded when NMR expression is close to IPSC expression and when NMR/IPSC expression is far away from human fibroblast expression. ChatGPT was used extensively throughout this project.
data and code
The comparison will be made with RNA sequencing data and comes from these links:
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE239446
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE294121
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE147871
The code which already includes the data is found here:
https://github.com/luojxxx/Comparing-gene-expression-for-aging
Results
Here’s a link to interpreting the results: https://docs.gsea-msigdb.org/#GSEA/GSEA_User_Guide/#interpreting-gsea-results
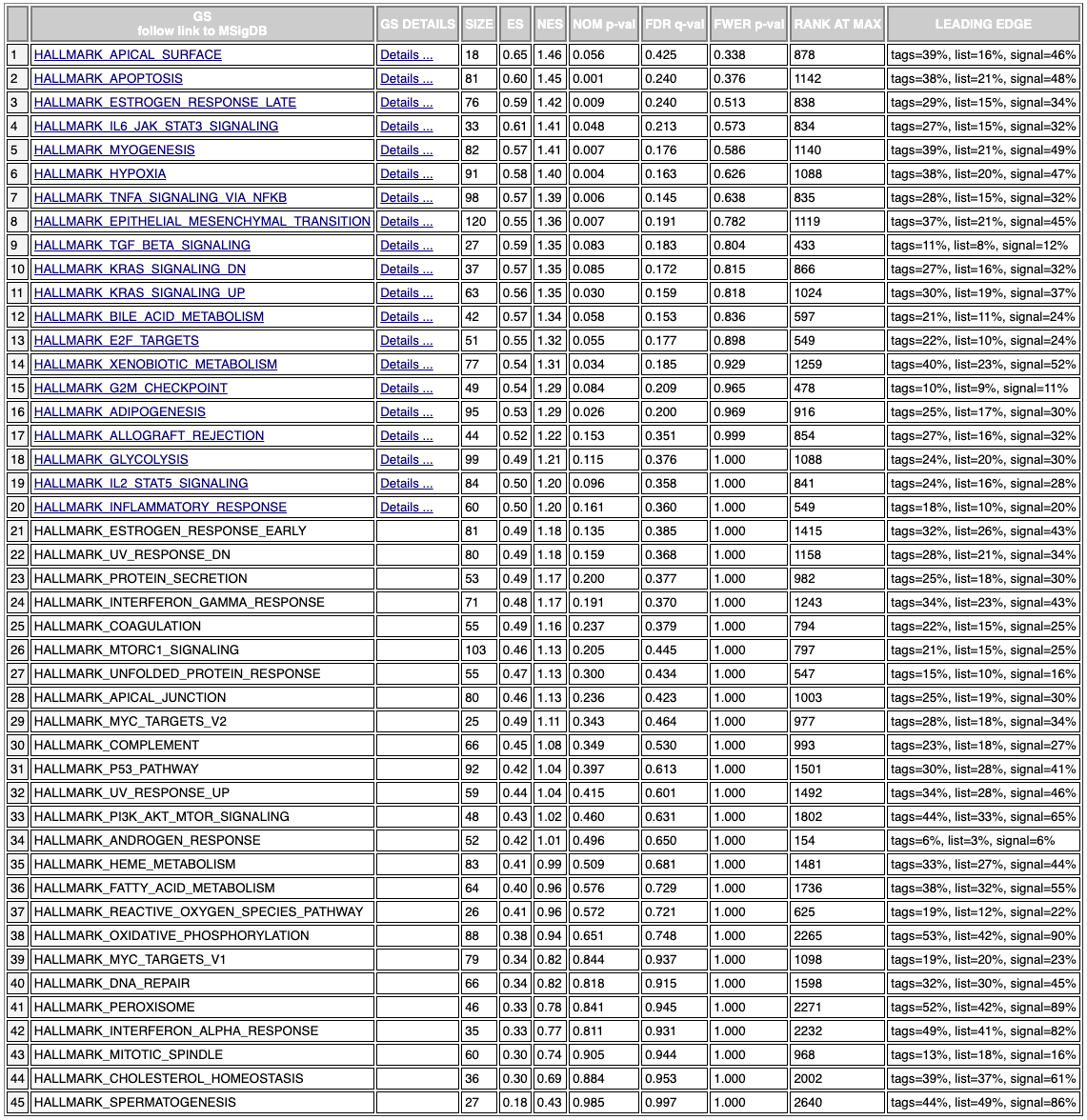
So after ranking the genes using my metric, we can see that apoptosis, inflammation, and cellular proliferation pathways for NMR fibroblast cells are similar to human IPSCs while different from human fibroblasts. It’s possible that NMR fibroblasts suppresses inflammation like how human IPSCs do, which is one of the reasons why they don’t age. Also the way the cells control apoptosis and cellular proliferation may be related to how the NMR avoids cancer and retains youthful cellular proliferation. Ultimately this exploratory study needs to be followed up with experimental studies to determine which genes have the most effect and how, although this exploratory study definitely focuses down on the list.
You can go through the results in detail under the metric run folder in the Github repo: https://github.com/luojxxx/Comparing-gene-expression-for-aging/tree/main/metric_run
The main results are:
https://github.com/luojxxx/Comparing-gene-expression-for-aging/blob/main/metric_run/metric_results.tsv
And the rnk file used for the gene set enrichment analysis is:
https://github.com/luojxxx/Comparing-gene-expression-for-aging/blob/main/metric_run/next_pass/gsea_no_confounds_agreement_and_stable_metric_eps_0p1.rnk
The outlier metric was something chatGPT came up with itself, but doesn’t follow what I wanted since it doesn’t reward closeness between the NMR fibroblast and human IPSC. The composite metric is the outlier metric divided by a denominator which ends up being my metric. Goes to show it still needs a human to watch the AI.